
Aleksandr Gaun
Software Engineer & Lab Automation Specialist
Engineer with a unique blend of life sciences expertise and software development skills. I've built high-throughput automation platforms, data analysis pipelines, and web applications across biotech companies including Calico Life Sciences and Expedition Medicines. I'm excited to apply my skills toward my vision for biotech lab automation and experiments. Published researcher with a patent in protein analysis methods. Now seeking opportunities to apply my cross-disciplinary background to software engineering, consulting, and freelance projects.
Resume/CV available on request — get in touch.
Projects

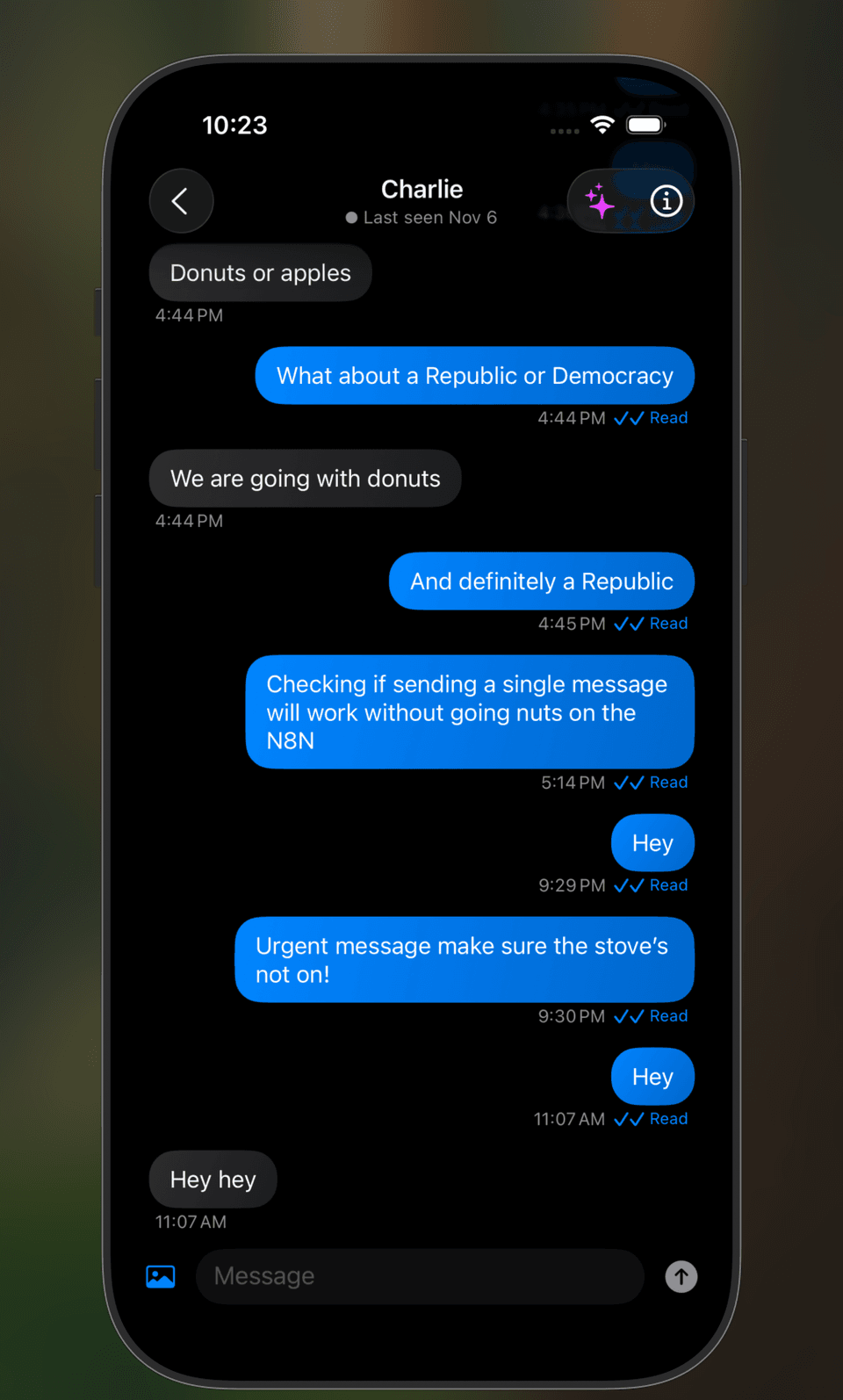
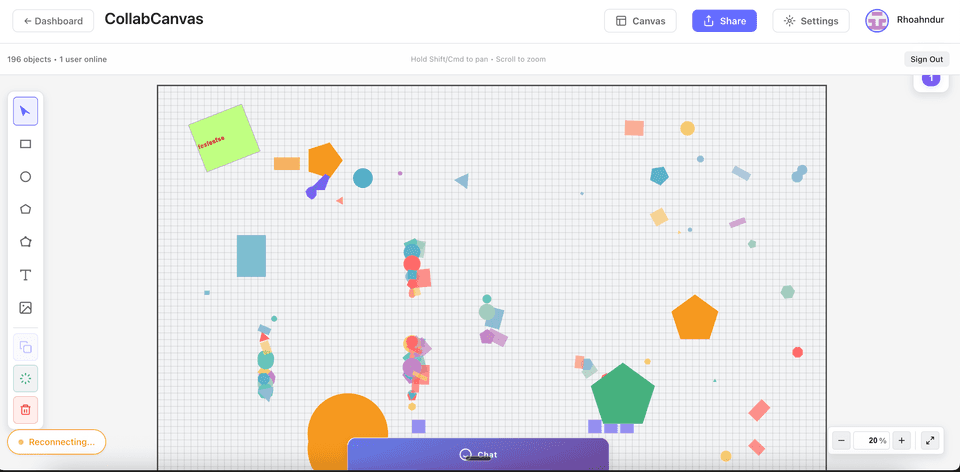
Smart iOS Messaging App
A modern iOS messaging application combining real-time chat with AI-powered features. Automatically extracts actionable insights, summaries, and action items from team conversations.
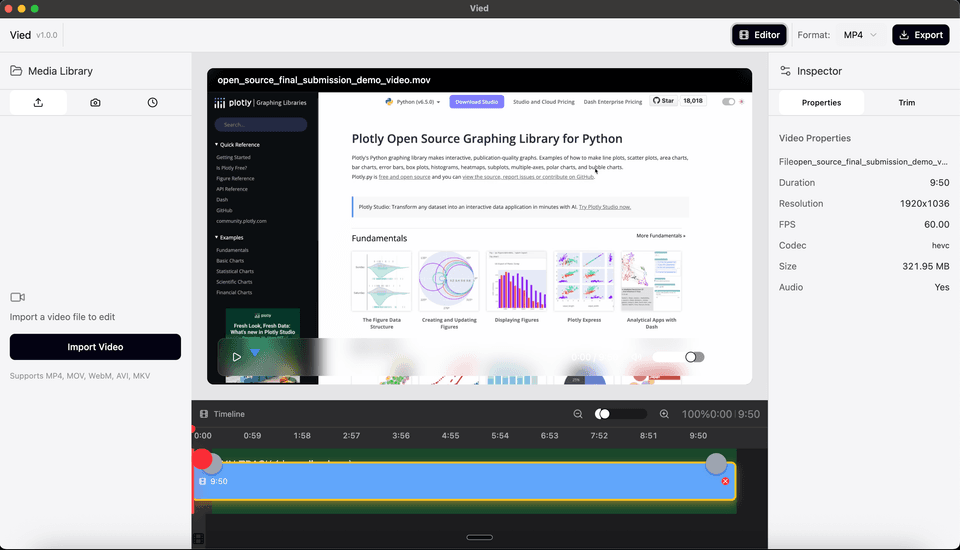
'Quick Cut' Video Editor
A desktop video editing application for trimming and exporting video files. Built with Electron and React, featuring a visual timeline with playhead indicator.
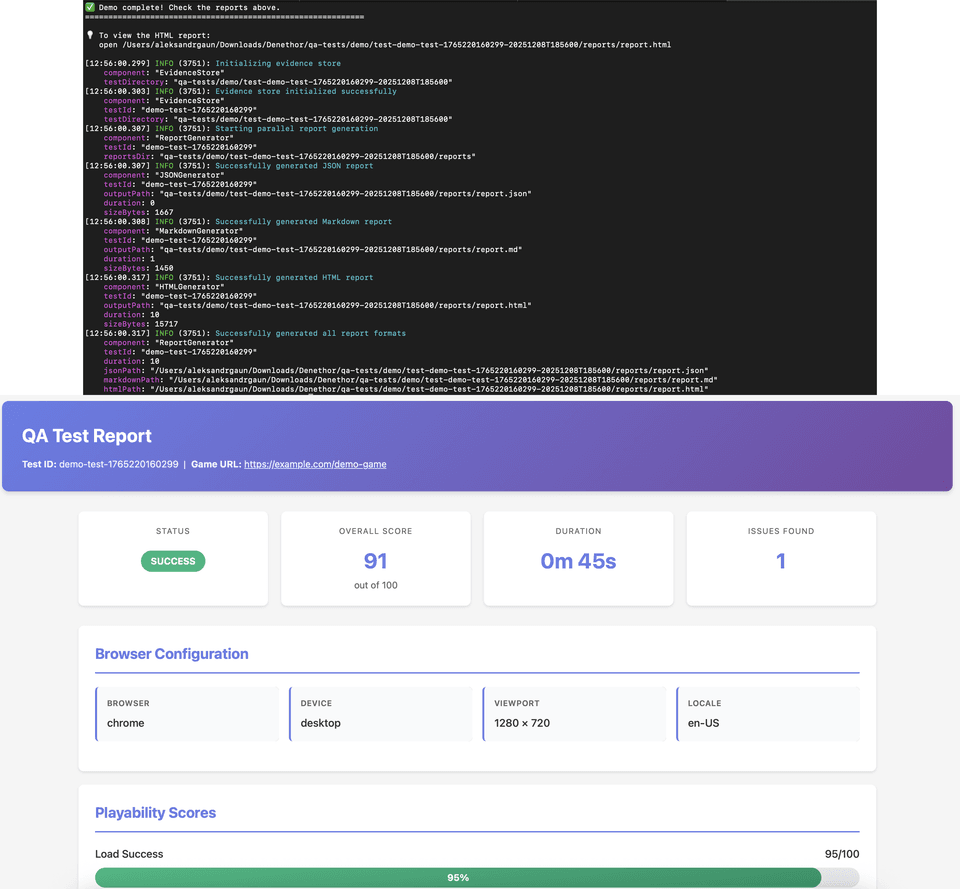
PlaytestBot
Autonomous AI agent for automated QA testing of browser-based games. Uses a hybrid approach combining heuristic patterns, vision-guided navigation, and GPT-4o evaluation.
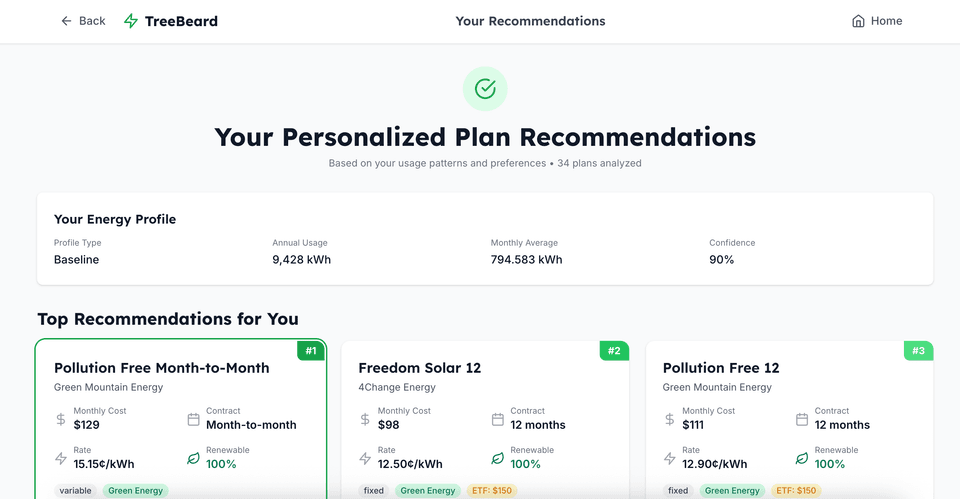
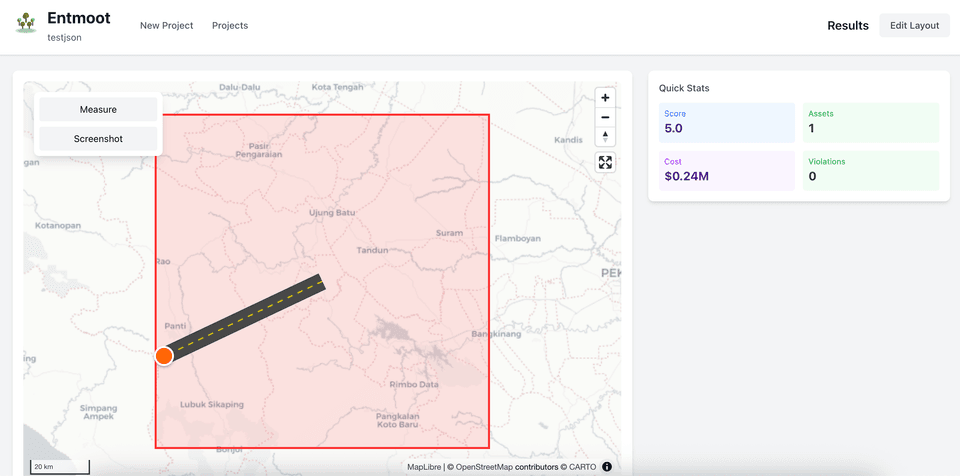
AI-powered energy plan recommendation system
AI-powered energy plan recommendation system for deregulated markets. Analyzes 12 months of usage data against 50+ plans to provide personalized recommendations with cost projections.
Open Source Contributions
Plotly.js
plotly/plotly.jsThe open source JavaScript graphing library that powers Plotly.
HumanLayer
humanlayer/humanlayerHuman-in-the-loop tooling for AI agents.
Experience
Senior Engineer II
May 2023 – Nov 2025Expedition Medicines · Cambridge, MA
Engineer / Principal Research Associate
Jan 2018 – May 2023Calico Life Sciences LLC · South San Francisco, CA
Research Associate
Jun 2014 – Jan 2018Harvard Medical School · Brookline, MA
Publications & Patents
Patents
- Methods of Protein Analysis. US20250110135
- Methods for analyzing proteomic attributes of biological samples and related systems and apparatus. WO2023239881
Publications
- Wang QS, Huang J, Chan LJG, Haste N, Olsson N, Gaun A, McAllister FE, Madhireddy D, Baruch A, Cardone KM, Kumar R, Ritchie M, Susztak K, Melamud E, Baryshnikova A. Platform-dependent effects of genetic variants on plasma APOL1. iScience (2026) doi:10.1016/j.isci.2026.114953
- Skarke C, Lahens NF, Mrčela A, Olsson N, Glessner J, Naik A, Theken KN, Chivily C, Douglas D, Hess KN, Gaun A, Moustafa AM, Daniel SG, Bittinger K, Teegarden S, Lordan R, El Jamal N, Meng H, Hollingsworth T, Woolfork A, Sengupta A, Brooks TG, Weljie A, Grant G, O'Brien J, McAllister F, Melamud E, FitzGerald GA. Age and the Diurnal Oscillatory Features of the Human Chronobiome. medRxiv (Preprint) (2026) doi:10.1101/2026.01.21.26344528
- Vu N, Maile TM, Gollapudi S, Gaun A, Seitzer P, O'Brien JJ, Hackett SR, Zavala-Solorio J, McAllister FE, Kolumam G, Keyser R, Bennett BD. Automated preparation of plasma lipids, metabolites, and proteins for LC/MS-based analysis of a high-fat diet in mice. Journal of Lipid Research (2024) doi:10.1016/j.jlr.2024.100607
- O'Brien JJ, Raj A, Gaun A, Waite A, Li W, Hendrickson DG, Olsson N, McAllister FE. A data analysis framework for combining multiple batches increases the power of isobaric proteomics experiments. Nature Methods (2024) doi:10.1038/s41592-023-02120-6
- Gaun A, Olsson N, Wang JCK, Eaton DL, McAllister FE. High-Throughput Proteome Profiling of Plasma and Native Plasma Complexes Using Native Chromatography. Methods in Molecular Biology (2023) doi:10.1007/978-1-0716-2978-9_5
- Gaun A, Preciado López M, Olsson N, Wang JCK, Chan LJG, O'Brien J, Li W, Zavala-Solorio J, Zhang C, Eaton D, McAllister FE. Triple-threat quantitative multiplexed plasma proteomics analysis on immune complex disease MRL-lpr mice. Proteomics (2022) doi:10.1002/pmic.202100242
- Gaun A, Lewis Hardell KN, Olsson N, O'Brien JJ, Gollapudi S, Smith M, McAlister G, Huguet R, Keyser R, Buffenstein R, McAllister FE. Automated 16-plex plasma proteomics with real-time search and ion mobility mass spectrometry enables large scale profiling in naked mole-rats and mice. Journal of Proteome Research (2021) doi:10.1021/acs.jproteome.0c00681
- Paulo JA, O'Connell JD, Gaun A, Gygi SP. Proteome-wide quantitative multiplexed profiling of protein expression: carbon-source dependency in Saccharomyces cerevisiae. Mol Biol Cell (2015) doi:10.1091/mbc.E15-07-0499
- Paulo JA, Gaun A, Gygi SP. Global Analysis of Protein Expression and Phosphorylation Levels in Nicotine-Treated Pancreatic Stellate Cells. J Proteome Res (2015) doi:10.1021/acs.jproteome.5b00398
- McDowell GS, Gaun A, Steen H. iFASP: Combining Isobaric Mass Tagging with Filter-Aided Sample Preparation. Journal of Proteome Research (2013) doi:10.1021/pr400032m
- Paulo JA, Gaun A, Kadiyala V, Ghoulidi A, Banks PA, Conwell DL, Steen H. Subcellular Fractionation Enhances Proteome Coverage of Pancreatic Duct Cells. BBA – Proteins and Proteomics (2013) doi:10.1016/j.bbapap.2013.01.011
- Paulo JA, Kadiyala V, Gaun A, Sauld JFK, Ghoulidi A, Banks PA, Steen H, Conwell DL. Analysis of endoscopic pancreatic function test (ePFT)-collected pancreatic fluid proteins precipitated via ultracentrifugation. Journal of the Pancreas (2013) doi:10.6092/1590-8577/1272
Education & Certifications
Education
Master of Science – Biochemistry & Molecular Biology
Clark University · Worcester, MA · 2014
Certifications & Training
- Hamilton Vantage Liquid Handler Venus 6 Advanced Sep 2024Hamilton Company, Reno, NV
- Hamilton Vantage Liquid Handler Venus 6 Beginner Aug 2024Hamilton Company, Reno, NV
- ASMS Short Course – Machine Learning for Mass Spectrometry Data Analysis Jun 2022Minneapolis, MN
- Hamilton Vantage Liquid Handler Instinct V Dec 2019Hamilton Company, Reno, NV
- Mini-MBA Certificate Jul 2016Harvard GSAS Business Club, Cambridge, MA